First, thanks so very much for the prompt reply.
and even more so for the great solution …
based on your example i used the following to prepare the pd correlation ouput:
mask = np.zeros_like(corr, dtype = np.bool) #probably could do this on df… to check
mask[np.triu_indices_from(mask)] = True
corr1=corr.mask(mask)
Then plot:
X = corr1.columns.values
sns_colorscale = [[0.0, ‘#3f7f93’],
[0.071, ‘#5890a1’],
[0.143, ‘#72a1b0’],
[0.214, ‘#8cb3bf’],
[0.286, ‘#a7c5cf’],
[0.357, ‘#c0d6dd’],
[0.429, ‘#dae8ec’],
[0.5, ‘#f2f2f2’],
[0.571, ‘#f7d7d9’],
[0.643, ‘#f2bcc0’],
[0.714, ‘#eda3a9’],
[0.786, ‘#e8888f’],
[0.857, ‘#e36e76’],
[0.929, ‘#de535e’],
[1.0, ‘#d93a46’]]
heat = go.Heatmap(z=corr1,
x=X, #check direct df column use
y=X, #check direct df row/index use
xgap=1, ygap=1,
colorscale=sns_colorscale, #“Magma”
colorbar_thickness=20,
colorbar_ticklen=3,
zmid=0 #added
hovertext =hovertext,
hoverinfo=‘text’
)
title = ‘Correlation Matrix’
layout = go.Layout(title_text=title, title_x=0.5,
width=800, height=800,
xaxis_showgrid=False,
yaxis_showgrid=False,
yaxis_autorange=‘reversed’)
fig=go.Figure(data=[heat], layout=layout)
fig.show()
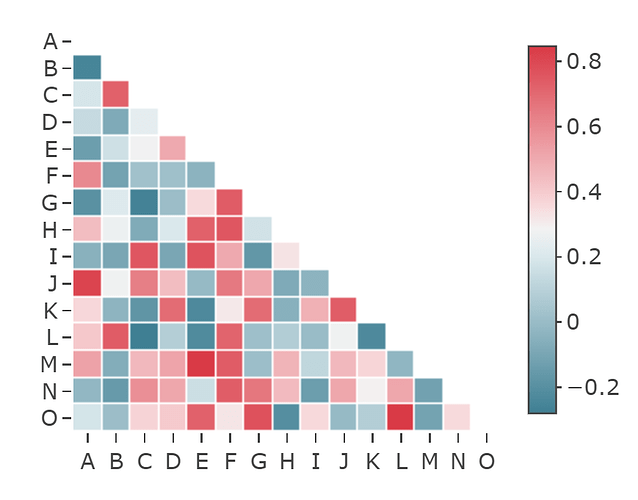
producing desired output:
one additional quick question…
what are these lines for? :
import plotly.io as pio
pio.templates.default = “none”
again many thanks
Rom
P.S.
really enjoying exploring how to use plotly